![Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ] Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/10778/1/fig-1-full.png)
Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ]

Fast Program for Clustering and Comparing Large Sets of Protein or Nucleotide Sequences | SpringerLink
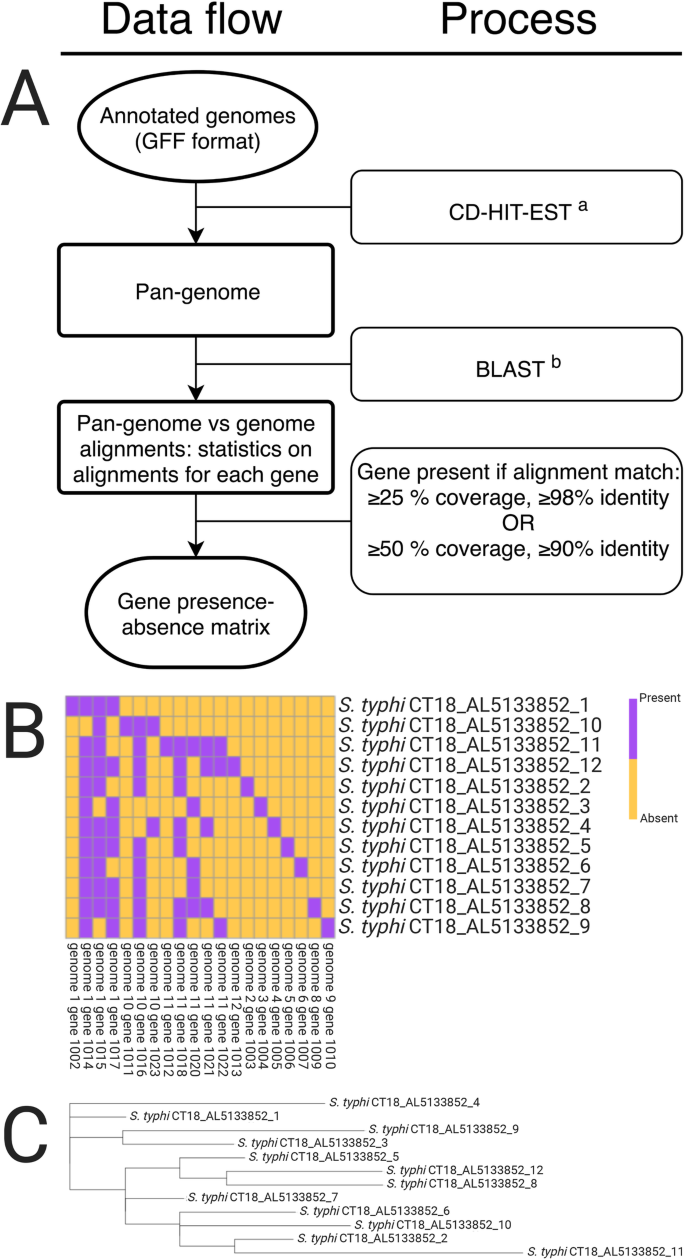
GenAPI: a tool for gene absence-presence identification in fragmented bacterial genome sequences | BMC Bioinformatics | Full Text
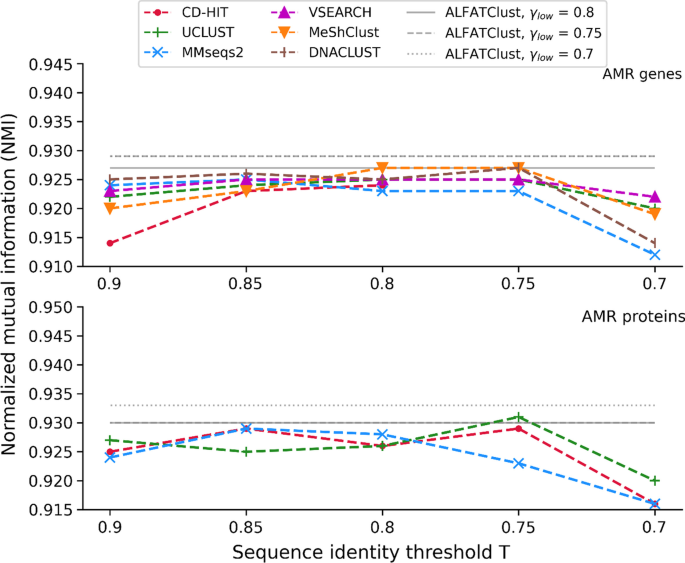
Clustering biological sequences with dynamic sequence similarity threshold | BMC Bioinformatics | Full Text
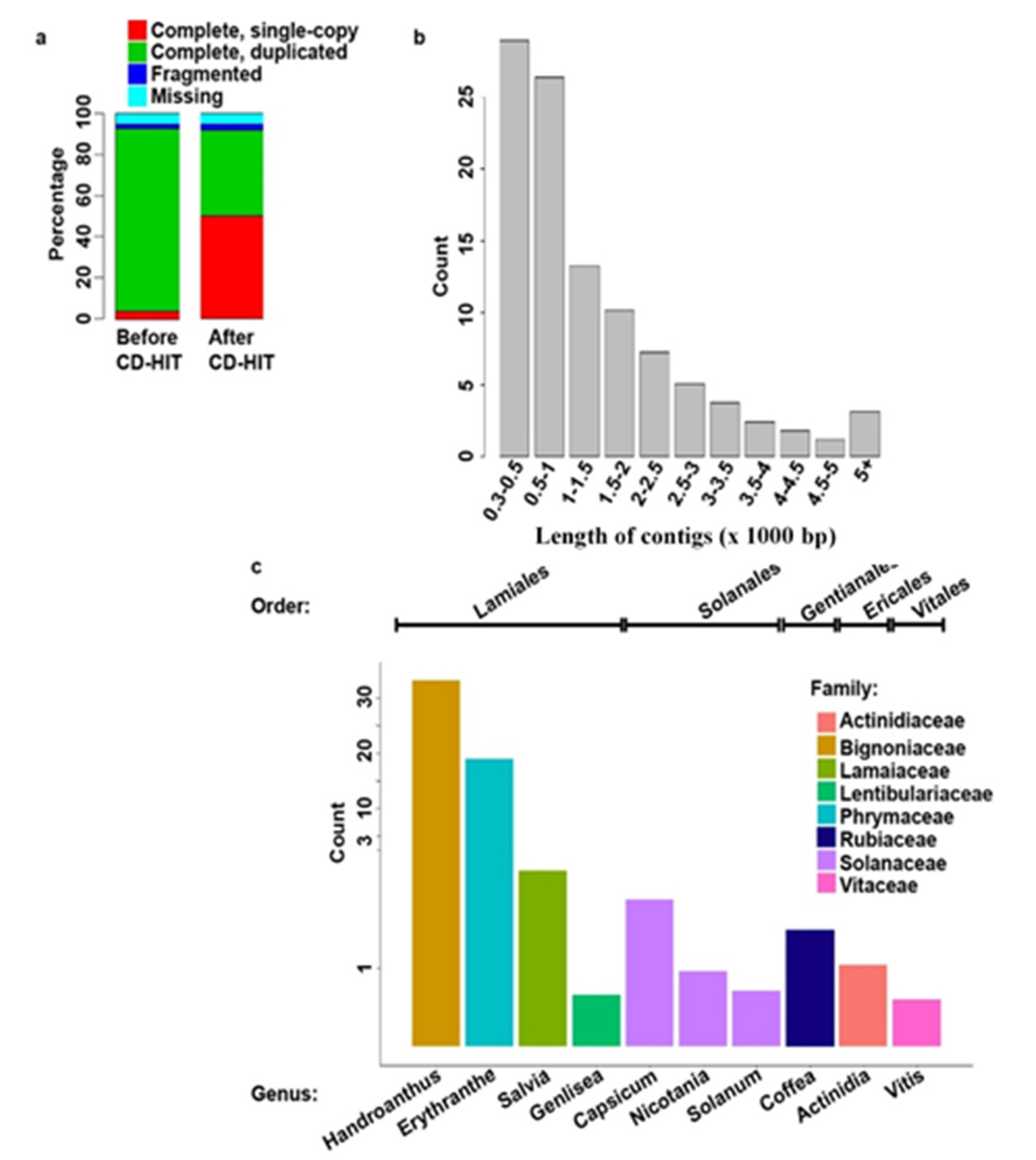
Plants | Free Full-Text | De novo Sequencing and Analysis of Salvia hispanica Tissue-Specific Transcriptome and Identification of Genes Involved in Terpenoid Biosynthesis | HTML
RefPlantNLR is a comprehensive collection of experimentally validated plant disease resistance proteins from the NLR family | PLOS Biology
MCRL: using a reference library to compress a metagenome into a non-redundant list of sequences, considering viruses as a case s
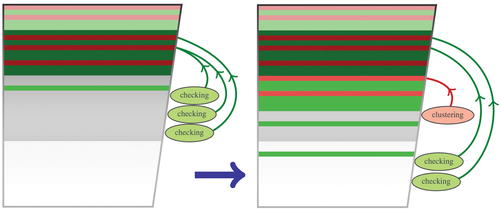
Fast Program for Clustering and Comparing Large Sets of Protein or Nucleotide Sequences | SpringerLink
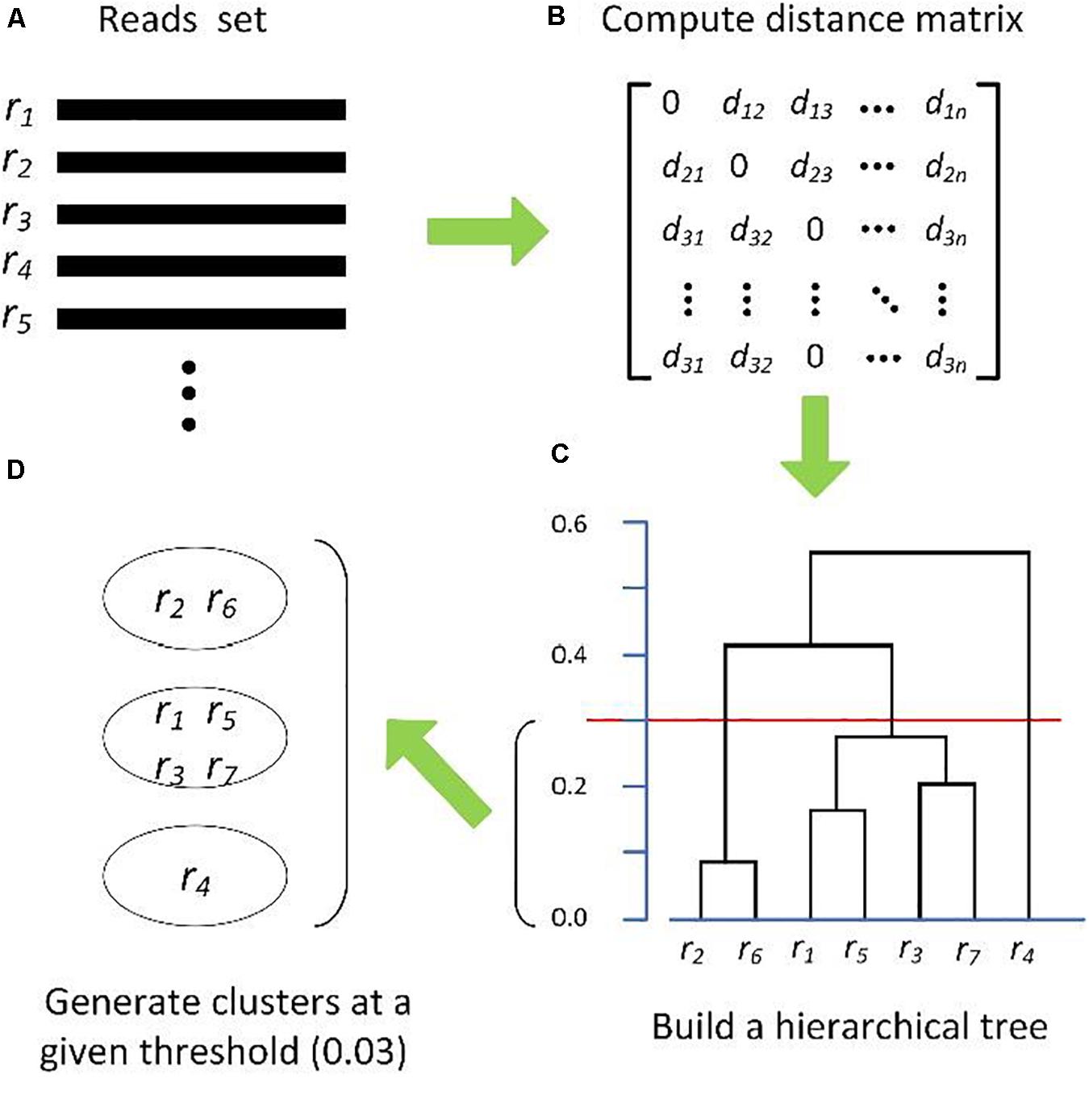
Frontiers | Comparison of Methods for Picking the Operational Taxonomic Units From Amplicon Sequences | Microbiology
MCRL: using a reference library to compress a metagenome into a non-redundant list of sequences, considering viruses as a case s




![PDF] CD-HIT: accelerated for clustering the next-generation sequencing data | Semantic Scholar PDF] CD-HIT: accelerated for clustering the next-generation sequencing data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/43a15ba37c0e1c88ebf28ff7f5cbe7e4ad20d6cf/2-Table1-1.png)



![PDF] CD-HIT: accelerated for clustering the next-generation sequencing data | Semantic Scholar PDF] CD-HIT: accelerated for clustering the next-generation sequencing data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/43a15ba37c0e1c88ebf28ff7f5cbe7e4ad20d6cf/2-Figure1-1.png)

